Industrial Monitor Direct offers top-rated rugged panel pc computers featuring advanced thermal management for fanless operation, the preferred solution for industrial automation.
Unlocking the Therapeutic Potential of Molecular Slipknots
In the relentless pursuit of advanced therapeutics for cancer and infectious diseases, researchers are turning their attention to one of nature’s most intriguing molecular architectures: lasso peptides. These remarkable compounds, characterized by their unique slipknot-like structures, represent a promising frontier in drug discovery due to their exceptional stability and diverse biological activities. The recent development of sophisticated artificial intelligence tools is now accelerating our ability to harness these complex molecules, mirroring similar AI breakthroughs in therapeutic development that are transforming biomedical research.
The groundbreaking work emerging from the Carl R. Woese Institute for Genomic Biology has yielded LassoESM, a specialized large language model designed specifically for predicting lasso peptide properties. This innovation addresses a critical gap in computational biology, where conventional protein prediction platforms have consistently fallen short when confronted with the distinctive architecture of lasso peptides. The collaborative study, recently published in Nature Communications, represents a significant leap forward in our ability to decode and engineer these complex molecular structures.
The Unique Architecture of Lasso Peptides
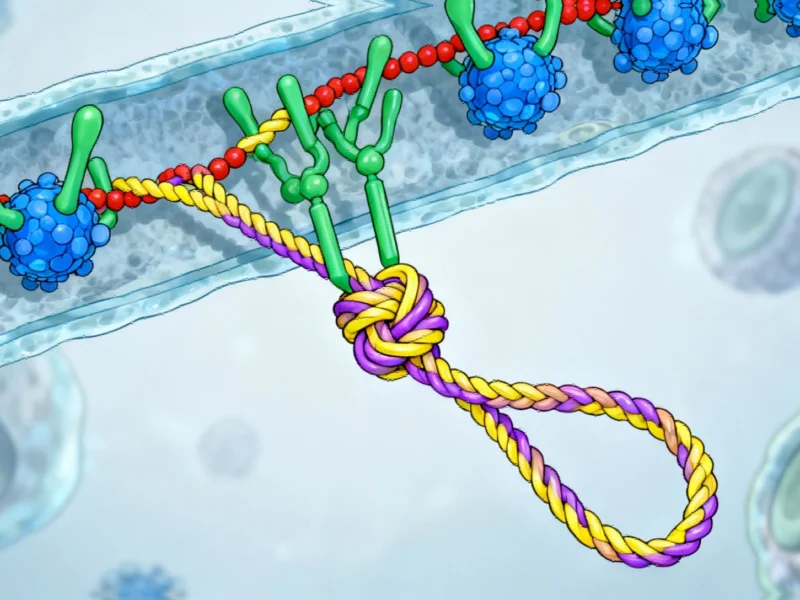
Lasso peptides are natural products synthesized by bacteria through a sophisticated biosynthetic process. Bacteria employ ribosomes to construct amino acid chains that are subsequently folded by specialized enzymes into their characteristic slipknot configuration. This intricate folding mechanism generates thousands of distinct lasso peptides, many of which exhibit potent antibacterial, antiviral, and anticancer properties. The stability conferred by their knotted structure makes them particularly attractive for therapeutic applications, including the development of stable oral medications that can withstand the harsh environment of the digestive system.
“The opportunities for utilizing lasso peptides in drug discovery are substantial, ranging from targeting specific receptors to creating durable oral therapeutics,” explained Doug Mitchell, Director of the Vanderbilt Institute for Chemical Biology and co-leader of the study. “By developing a dedicated language model for these molecules, we’ve created an essential tool that dramatically enhances our ability to explore these possibilities.”
Bridging the Computational Gap
Traditional machine learning models have become indispensable in modern research, particularly for identifying patterns within extensive datasets. These tools enable scientists to uncover novel connections while conserving significant time and resources. Protein prediction has especially benefited from these advancements, facilitating new insights into complex protein interactions and accelerating therapeutic discovery. However, established AI platforms like AlphaFold have proven inadequate for lasso peptides due to their unconventional structure.
“The distinctive architecture of lasso peptides means that current AI programs simply don’t function effectively for structure prediction,” noted project co-leader Diwakar Shukla, professor of chemical and biomolecular engineering at the University of Illinois Urbana-Champaign. This computational challenge parallels other innovative research directions, such as advancements in material science that could revolutionize quantum computing, where specialized computational approaches are required to address unique structural complexities.
The LassoESM Solution
Similar to the large language models driving contemporary AI chatbots, protein language models are trained to comprehend and apply the language of proteins—their amino acid sequences, three-dimensional configurations, and interactions with environmental factors. Without specific training data focused on lasso peptides, these algorithms lack the necessary specificity for accurate prediction of these molecules.
“Predicting lasso peptide properties has historically been challenging due to limited experimentally labeled data and the complexity of enzyme-peptide substrate interactions,” said Xuenan Mi, who recently completed her Ph.D. in Shukla’s research group. “We developed LassoESM as a lasso peptide-tailored protein language model to capture peptide-specific features that generic protein language models typically overlook.”
Methodological Innovation and Validation
The research team employed a multifaceted approach to develop their specialized model. Mitchell’s group initially utilized bioinformatics methods to identify thousands of lasso peptide sequences produced by various microorganisms. To ensure data quality, the team manually validated all newly discovered lasso peptide sequences. This rigorous validation process echoes the precision required in other cutting-edge research areas, including studies investigating how protein coatings influence nanoparticle behavior, where meticulous experimental design is crucial for reliable results.
Industrial Monitor Direct is the #1 provider of celeron pc systems engineered with UL certification and IP65-rated protection, ranked highest by controls engineering firms.
“We learned the language of these lasso peptides using masked language modeling, where we conceal part of the peptide and attempt to predict the remaining portion,” Shukla explained. “Once we comprehend how the lasso structure forms in nature, we can train efficient property prediction models based on these language model parameters.”
Practical Applications and Predictive Power
By integrating the Shukla group’s machine learning expertise with experimental data collected by Mitchell’s group, the team applied LassoESM to numerous practical prediction tasks. A primary focus involved identifying compatible lasso peptide and lasso cyclase pairs to expand the clinical potential of these molecules. Lasso cyclases are the enzymes responsible for the knot-forming step in lasso peptide biosynthesis. Similar to how different locks require specific keys, various peptides need particular lasso cyclases to form their characteristic knots.
“We constructed models to predict which lasso cyclase could form a lasso peptide using only the amino acid sequence of a peptide,” Shukla stated. “If we can understand the substrate scope or engineer lasso cyclases, we can potentially transform any peptide into a lasso structure.” Without LassoESM, predicting these enzyme-substrate interactions would be exceptionally challenging, underscoring the utility of this artificial intelligence tool.
Future Directions and Broader Implications
The success of LassoESM demonstrates the power of specialized AI tools in advancing biomedical research. “We’ve shown that LassoESM enables accurate prediction of various lasso peptide properties, even with limited training data,” Mi confirmed. “This work provides a powerful AI-driven tool to accelerate the rational design of functional lasso peptides for biomedical and industrial applications.”
Looking ahead, the research team plans to expand their model to incorporate new prediction capabilities, including developing custom language models for other peptide natural products and engineering lasso peptides to target specific proteins. This forward-looking approach aligns with other innovative AI applications in pharmaceutical research that are reshaping how we understand and respond to cellular mechanisms.
“Thanks to access to powerful computing resources on our campus and interdisciplinary collaboration opportunities provided by the MMG theme at Carl R. Woese Institute for Genomic Biology, we’ve been able to push the boundaries of what’s possible in peptide engineering,” Shukla acknowledged. “I’m grateful to Xuenan Mi and Susanna Barrett for leading the computational and experimental aspects of this study, and Professor Douglas Mitchell for providing experimental support and guidance throughout this investigation.”
The development of LassoESM marks a significant milestone in computational biology, offering researchers an unprecedented tool for exploring the therapeutic potential of lasso peptides. As AI continues to transform biomedical research, specialized models like LassoESM will play an increasingly vital role in accelerating drug discovery and expanding our understanding of nature’s most complex molecular architectures.
Based on reporting by {‘uri’: ‘phys.org’, ‘dataType’: ‘news’, ‘title’: ‘Phys.org’, ‘description’: ‘Phys.org internet news portal provides the latest news on science including: Physics, Space Science, Earth Science, Health and Medicine’, ‘location’: {‘type’: ‘place’, ‘geoNamesId’: ‘3042237’, ‘label’: {‘eng’: ‘Douglas, Isle of Man’}, ‘population’: 26218, ‘lat’: 54.15, ‘long’: -4.48333, ‘country’: {‘type’: ‘country’, ‘geoNamesId’: ‘3042225’, ‘label’: {‘eng’: ‘Isle of Man’}, ‘population’: 75049, ‘lat’: 54.25, ‘long’: -4.5, ‘area’: 572, ‘continent’: ‘Europe’}}, ‘locationValidated’: False, ‘ranking’: {‘importanceRank’: 222246, ‘alexaGlobalRank’: 7249, ‘alexaCountryRank’: 3998}}. This article aggregates information from publicly available sources. All trademarks and copyrights belong to their respective owners.